Difference between revisions of "VolCT - Group 1A"
| Line 102: | Line 102: | ||
'''Powerpoint examples of nodules with Filters''' | '''Powerpoint examples of nodules with Filters''' | ||
| − | [[Media: | + | [[Media:QIBA-slicedata-medium-detailed-2009-02-02.ppt|2009-02-02: Examples of Philips Medium and Detailed reconstruction filter slices-]] |
| + | ]] | ||
Revision as of 20:51, 6 February 2009
Current Goals & Status:
- Goal: Benchmark intra/inter-reader variability for lung nodule volume measurements
- Status: Most of Dataset acquired; finalizing reader software
- Related Profile: Lung Vol Quantification - Measurement Activity
Work Documents
- Vol-CT - 1A Group Call Summaries
- Proposed 1A Protocol Outline (2008-11-13-v05)- For Comment from QIBA Members
- ---- Comments Bob Ford 12/16/2008
- RSNA Abstract SSK04-04: phantom study simialr to our proposal- For discussion on 12/18/2008 1A t-con.
- 2002-01-26: 1A Reader Study timeline for completion 1/1-4/15/2009- Please edit/comment
- 2009-02-02: Examples of Philips Medium and Detailed reconstruction filter slices- Please review/vote on appropriate filtering for 1A study
- 2009-02-02: 1A study 40 nodule slice date- 10mm ellipse -> 20 mm ellipse
Projects
May want to split these out to separate pages later
VolCT Lung Anthropomorphic Phantom Study
Objective:
Measure intra- & inter-reader bias and variability phantom lesions for:
- Uni-dimensional size measurement
- Semi-automatic 3D volumetric measure
May compare with fully automated algorithm(s)
Dataset:
Ground truth has been established by physical measurement "ex vivo“ on FDA phantom inserts.
Nodules (10 attached nodules)
- -10 & +100HU
- 10, 20 mm spheres
- 10 mm ovoid, lobulated, spiculated
Image Dataset
- 100 mAs exposures
- 0.75 & 5.0 mm slices
- 1 recon kernel
- Status:
- mostly acquired by FDA/CDRH/OSEL.
- Missing: 10 mm ovoid, spic, lob (Est: 12/01/08)
Acquisition Protocol:
- Scanner: Philips 16 slice
- Exposure (120 kVp): 100 mAs
- Slice thickness (50% overlap): 0.75 & 5.0 mm slices
- Recon kernel: Standard/medium (Still on table: Detail/Lung kernel)
- Pitch: 1.2
- 2 repeat scans
- 40 segmentations in set
The study is being conducted as a pilot. The size (data, readers) has not been selected for any specific level of significance.
Study Protocol:
Expert readers will measure/estimate nodule size from CT images.
Readers: 6 RadPharm radiologists
Software:
- Wendy will visit RadPharm and provide more info next week
- In-house review software (Siemens?)
- Semi-automated 3D volume software
- Uni-dimensional measure (RESIST)
- Which fully automated software?
Reading Session:
- Readers read all cases in 2 different reading sessions
- Random ordering
- One-dimensional measure
- Semi-automated segmentation
- Sessions separated by 3 week(?)
- Include duplicate cases within each read session (1/3-1/2 of cases for intra-reader estimates)
- Random ordering
- Time restrictions: Probably not (?)
- Specific instructions: Probably not (?)
- Readers read all cases in 2 different reading sessions
Analysis:
Estimate intra- and inter-reader variability in the different volume estimate
- Estimate bias from known truth
- Estimate variability
Compare the bias and variability with the different methods
Outstanding Issues:
1A Questions to Group
Please selection either Medium or Detail Reconstruction Filter for 1A reader study
Philips Filter Descriptions
Detail: Medium-sharp filter, defined either for medium resolution images or moderate-noisy conditions of acquisitions (such as Posterior Fossa, chest and abdomen)
Medium: defined either to enhance the low contrast (for example Posterior Fossa) or to decrease the noise when the acquisition has been performed in relatively noisy conditions (such as large patient, thin slices and low mAs)
Examples of Detailed (Left) and Medium (right) filters
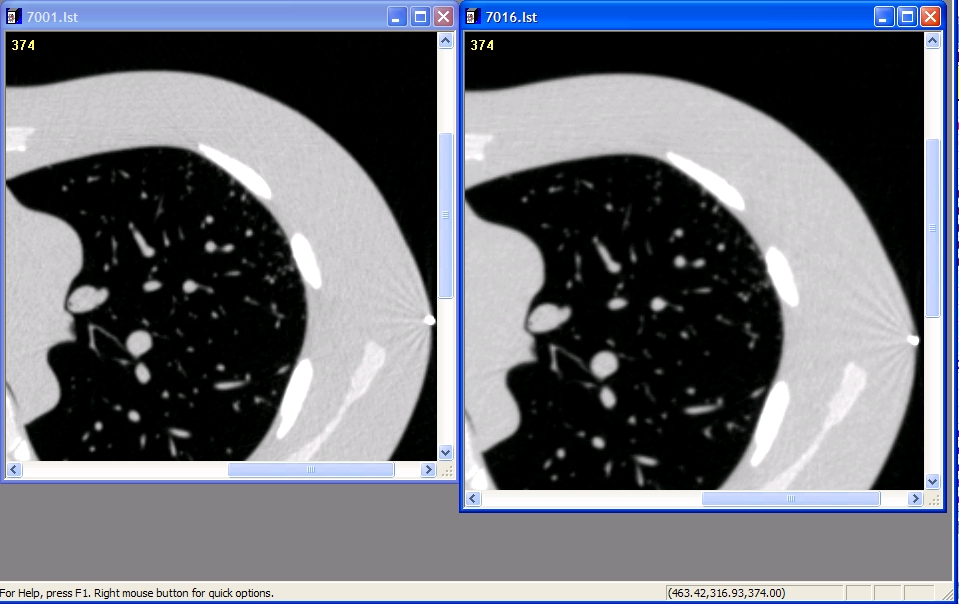
Powerpoint examples of nodules with Filters 2009-02-02: Examples of Philips Medium and Detailed reconstruction filter slices- ]]

